 The tool creates images of 2D-spots for the evaluation and benchmarking of spot detection tools. 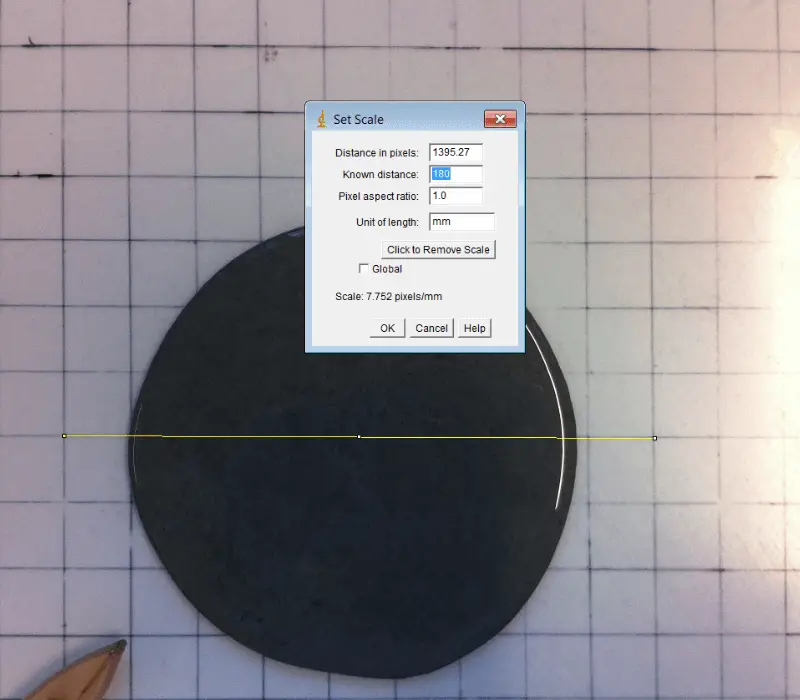
The tool measures the mean distance between the minima in the profile plot. The tool uses expectation maximisation clustering from the weka software. The populations are separated by the area of the spots. It separates the spots into two populations and counts each population individually. It has been created for the counting of bacteria colonies in in Petri-dishes. The tool detects and and counts the spots (or blobs) in an image. The tool detects and counts the neurons and the neurons with satellite cells. The tool detects and counts the maxima along a line-selection, for example a segmented line. Count Axonal Projections ToolĬount the number of axonal projections that cross a given line. The tool allows to make the MIP-projections of the two kinds of input images in batch mode. After the detection of the spots the background has been set to 0 and the volumes of the spots to 255 and the image has been exported as a tif-series. The tool needs two types of input images: the 3d stack of the hair cells and a binary mask created from this stack by using the spot detection algorithm of Imaris (Bitplane). The aim of this tool is to count the hair cells in sections of 200µm from the apex of the cochlear to its base. The nearest neighbor distances between all nuclei, and those outside and inside of the clusters are calculated. The resulting points are clustered using the DBSCAN algorithm. Using a threshold intensity value, maxima below the threshold are eliminated. The nuclei are detected as the maxima in the image. Cluster Analysis of Nuclei ToolĪnalyze the clustering behavior of nuclei in DAPI stained images. It writes a spreadsheet file with the measured area per well and saves a control image showing the green surface that has been detected per well. It can be run in batch mode on a series of images. The Arabidopsis Seedlings Tool allows to measure the surface of green pixels per well in images containing multiple wells. If a third channel is provided it is used to filter out detected protoplasts that do not have exactly one nucleus. 
The tool counts the spots per protoplast. It can also count and measure the area of the nuclei within the speroïd. The tool allows to measure the area of the invading spheroïd in a 3D cell invasion assay. Analyze Spheroid Cell Invasion In 3D Matrix Instead of circles, which present the distance from the base point, horizontal lines can be used, which present the distance in the soil from the base-line. It then allows to draw concentric circles around a base point and to extract measures on or within the circles. The tool creates a binary mask and the Euclidean Distance Transform from the input image. The tool is inspired by the Sholl analysis used in neuronal studies. While for young roots the root system architecture can be analyzed automatically, this is often not possible for more developed roots. This tool allows to analyze morphological characteristics of complex roots. 
The tool measures the areal number density of comets in cells. Although the input images can be stacks only the middle slice is used for the analysis. The tool measures the length of the sarcomeres using the FFT of the image and the degree of organization of the sarcomeres by using the dispersion provided by the Directonality command of FIJI. Analyze_CardiomyocytesĪnalyze images from second harmonics microscopy of cardiac muscle cells (cardiomyocytes). It measures the intensity of the calcium signal in the whole region of interest and in the segmented spots. The tool finds the region to analyze close to the point of stimulation. Each image contains a selection that marks the point of stimulation. The images consist of time-series of calcium signals. Analyze_Calcium_Signals_In_SpinesĪnalyze calcium signals in dendritic spines. The direction-histograms and the measurements are exported as csv-files. 
The tool allows to run the Directionality plugin in batch-mode on a series of images. It is used in this context to analyze to which degree the muscles in the image are vertically aligned. The tool uses the Directionality plugin to measure the main direction of the structures in the image and the dispersion. The Adipocytes Tools help to analyze fat cells in images from histological section. The nearest neighbor distances between all nuclei and those outside and inside of the clusters are calculated. The clustering is done with the density based algorithm DBSCAN. The nuclei are filtered by the presence of a signal in a different channel. Biological Image Analysis Toolsets 3D_Nuclei_Clustering_ToolĪnalyze the clustering behavior of nuclei in 3D images. Have a look at the project’s wiki to get more information about the image analysis tools. ImageJ macros and scripts written at the imaging facility MRI. Imagej_macros_and_scripts Imagej macros and scripts 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
